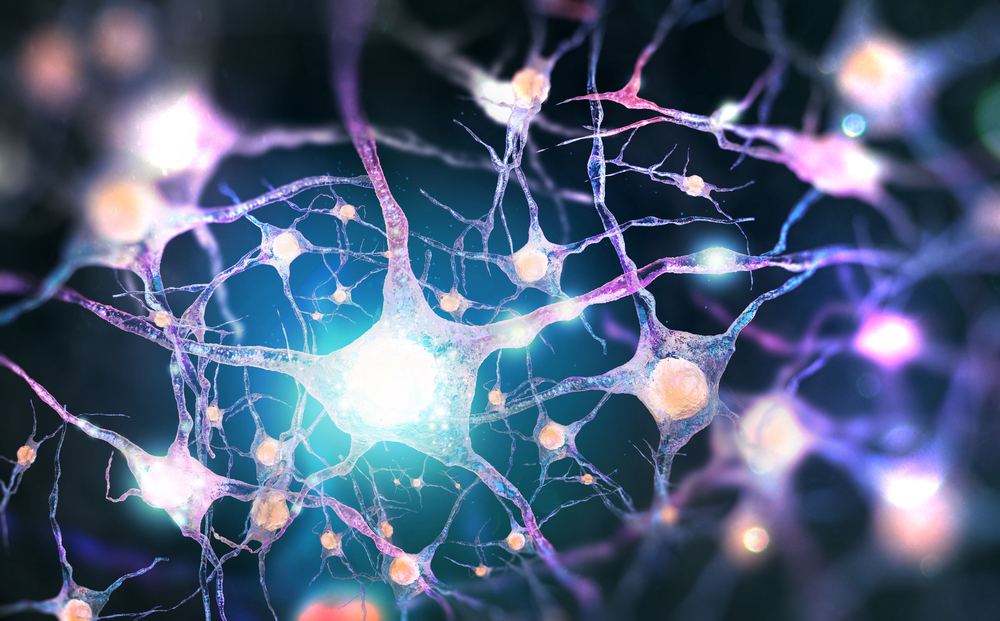
Sequencing reads of a sample prepared by the traditional whole-genome sequencing workflow for fresh-frozen samples and data generated using the SeqPlus whole-genome FFPE method. The green and purple indicate reads sequenced in the forward and reverse directions, respectively, and yellow represents bases with non-reference sequence. The center of the image shows a C to A mutation in each of the tumor samples.
Cancer is one of the most active fields in genomics, spurring mountains of research papers and scores of clinical trials. WuXi NextCODE (WXNC) is committed to pushing this field forward and so we had a special “Genomes for Breakfast” session devoted to this topic at the recent ASHG17 event. Featured talks addressed our pathbreaking work in how to extract impactful findings from the renowned TCGA dataset; get better sequencing results from FFPE samples; and apply deep learning to drug discovery, drug repurposing, and identifying subtypes for diagnostics and clinical trials.
The Cancer Genome Atlas (TCGA) is one of the most useful public genomic cancer databases available and has already led to numerous critical discoveries, including entirely new drug targets as well as better insights into tumor origination, development, and spread. It includes data from approximately 11,000 patients and covers 33 cancer types. Data types include WES, RNA-Seq, mi RNA, CNV, Methylation array, and clinical sample data. The data is big and complex, and can include multiple samples from one patient, which is crucial to know when doing analyses.
During his ASHG talk, Jim Lund, WXNC’s Director of Tumor Product Development, shared some insights into how we put this rich data source to work in concert with our own unique data and analytical tools, in a process he dubs “multiomics analysis.” He described how we specially process the data and use our unique analytical platform to help scientists find just what they are looking for. Researchers can search the data by cancer type, age of diagnosis, sex, ethnicity, year of diagnosis, sample type (e.g. metastatic, new primary), and more.
Multiple pivotal studies using this dataset have already been published, including some examining the prevalence of specific mutations across human cancer types as well as in-depth profiling of specific tumors, such as breast cancer and lung adenocarcinoma. Layering different types of data, such as reads from DNA and RNA, allows much more accurate detection of features such as variants with allele-specific effects on gene expression. The user-friendly but sophisticated data interface makes it easier to see such findings. Over the years, our own database and our capabilities have both grown exponentially, creating a powerful tool for multiomics cancer research. You can see Jim putting the portal through its paces in a recent webinar.
In his talk, Shannon Bailey described how Whole Genome Sequencing (WGS) can be applied to formalin-fixed paraffin-embedded (FFPE) tumor samples, which are stored by the hundreds of thousands in repositories around the world. Shannon is the Associate Director of our Cancer Genetics division. He pointed out that while these samples are abundant and often paired with extensive clinical and outcome data, there are specific hurdles to using these for the type of large-scale retrospective studies many groups are eager to carry out.
For one thing the genetic material in such samples can be degraded, crosslinked, or in low quantities. Of all these problems, the biggest issue is getting sufficient quantity of quality DNA for sequencing. Numerous studies have found that these types of samples are difficult to work with and often provide very low success rates for gene sequencing studies. Clearly, fresh frozen samples provide much better results, but they are also much harder to obtain.
In response, our team has developed the WXNC SeqPlus FFPE extraction method, which provides substantially improved coverage compared to traditional methods and even approximates the results obtained with fresh frozen samples at 10X depth, with similar numbers of heterozygous and homozygous calls.
We tested SeqPlus in a study that comprised 516 tumor-normal pairs (i.e., 1,032 samples) that had been stored for 3 to 6 years. The targeted sequencing depth was 30X for the normal tissue and 70X for tumor tissue. The starting amount of DNA was 400 ng. The results were excellent, with SeqPlus delivering a coverage analysis just about 1% below what the fresh frozen control samples achieved. Further, a comparison of our analyses to results from the TCGA, using fresh frozen samples, showed striking similarity. These study results give us confidence that SeqPlus is a new “power tool” for FFPE sequencing studies. This webinar describes the process.
Sequencing reads of a sample prepared by the traditional whole-genome sequencing workflow for fresh-frozen samples and data generated using the SeqPlus whole-genome FFPE method. The green and purple indicate reads sequenced in the forward and reverse directions, respectively, and yellow represents bases with non-reference sequence. The center of the image shows a C to A mutation in each of the tumor samples.
Another area of great interest at WXNC is artificial intelligence (AI). We have been pioneers in AI for pulling novel insights out of massive multiple datasets. Leading this effort is Tom Chittenden, our Vice President of Statistical Sciences, Founding Director of the Advanced AI Research Labs, and a Lecturer on Pediatrics and Biological Engineering at Harvard Medical School and MIT. He also spoke at the breakfast series.
Our AI capabilities improve the tools we have and expand their capabilities. For example, using our AI tools, we can improve functional annotation of missense variants to an accuracy of >99%, integrate multiple types of data to discover new genes and elaborate pathways, and improve tumor subtype and drug-response classification accuracy by combining DNA- and RNA-seq, among other data types. These tools can be used for such varied purposes as target discovery, drug repurposing, and defining responders and non-responders in clinical trials.
We’ve already helped to develop breakthrough results, such as identifying an intriguing new target for both cardiovascular and cancer drug discovery. We’ve also classified breast and lung cancer subtypes with 97% to 100% accuracy, classified 8,200 tumors of 22 TCGA cancer types with >99% accuracy, and discovered a completely novel pan-cancer molecular survival signature.
The power of our deepCODE AI tools is in part thanks to a novel, causal statistical-learning method and deep-learning classification strategy. But another advantage is that they were built on our global platform for genomic data, which underpins the majority of the world’s largest genomics efforts and includes all major global reference databases. Our database stores, manages, and integrates any type of genomic data and correlates it with phenotype, ‘omics’, biology, outcome, and virtually any other type of data that may be relevant to a particular medical challenge.
If you want to know more, I recently gave an interview to WXpress outlining WXNC’s AI strategy. As we continue to deepen our commitment to this field, I’m sure we’ll have more exciting results to share.